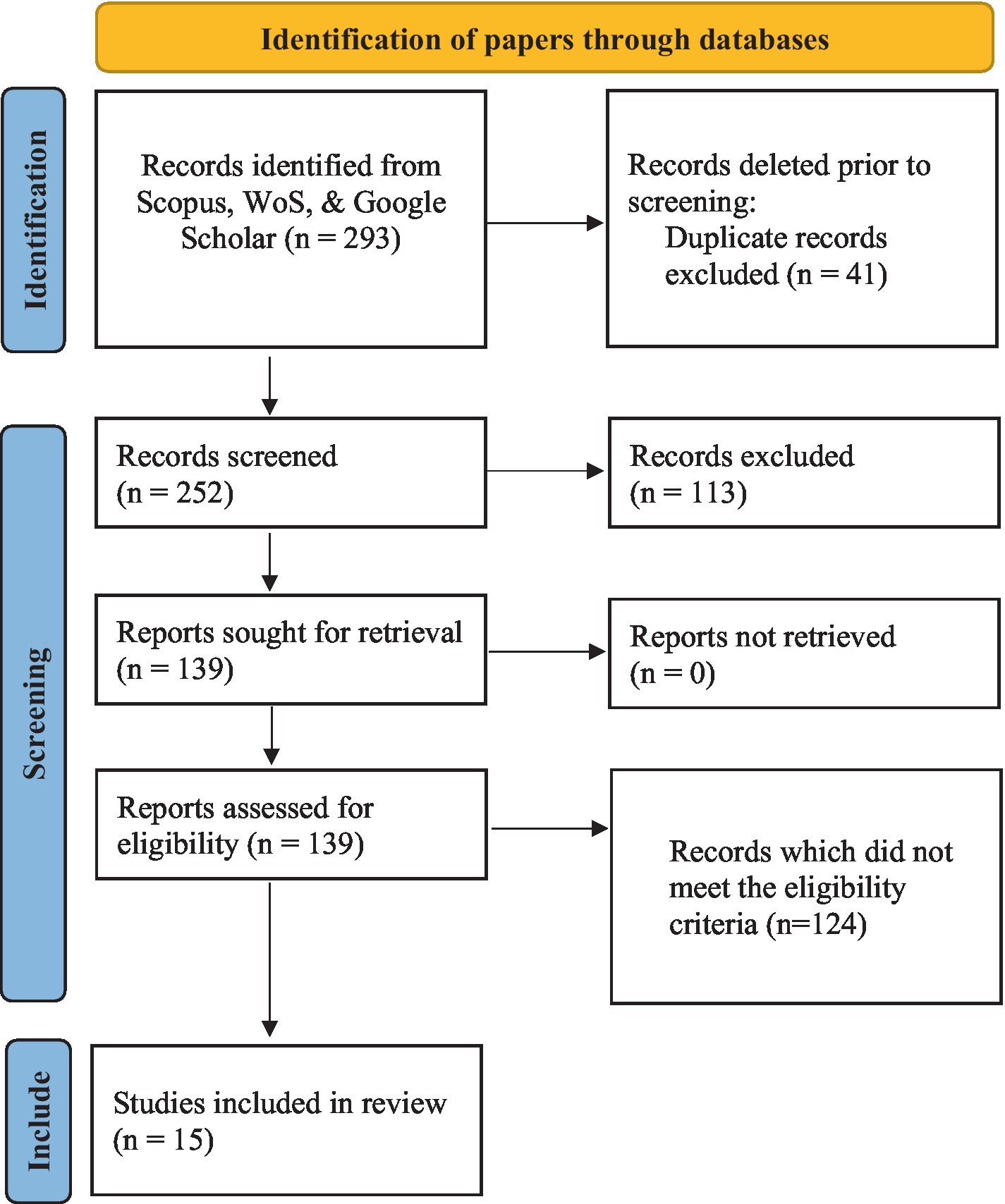
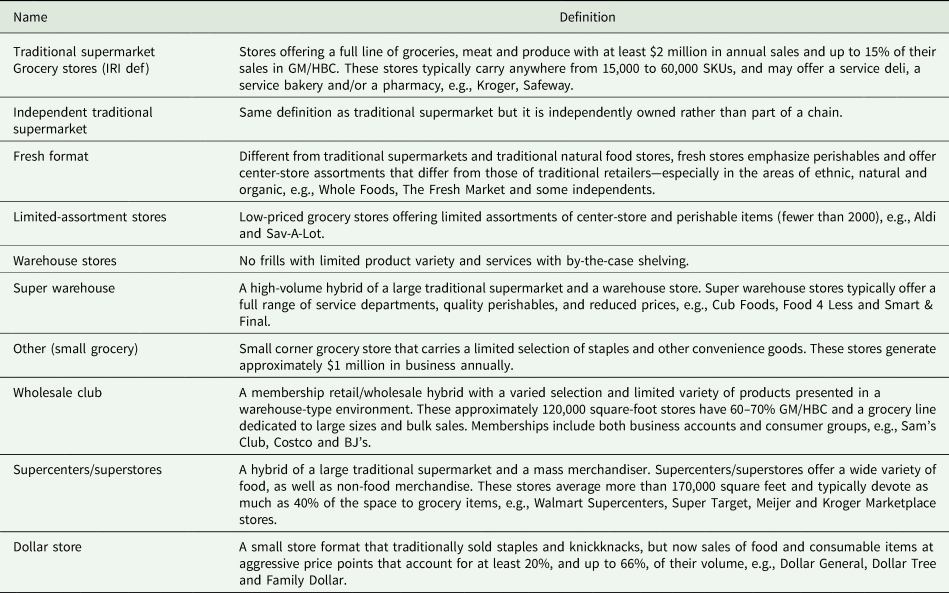
Structural basis for sequence-independent substrate selection by eukaryotic wobble base tRNA deaminase ADAT2/3
By A Mystery Man Writer
Description

Neurodevelopmental disorders associated variants in ADAT3 disrupt

Structural basis of tRNA recognition by the m3C-RNA
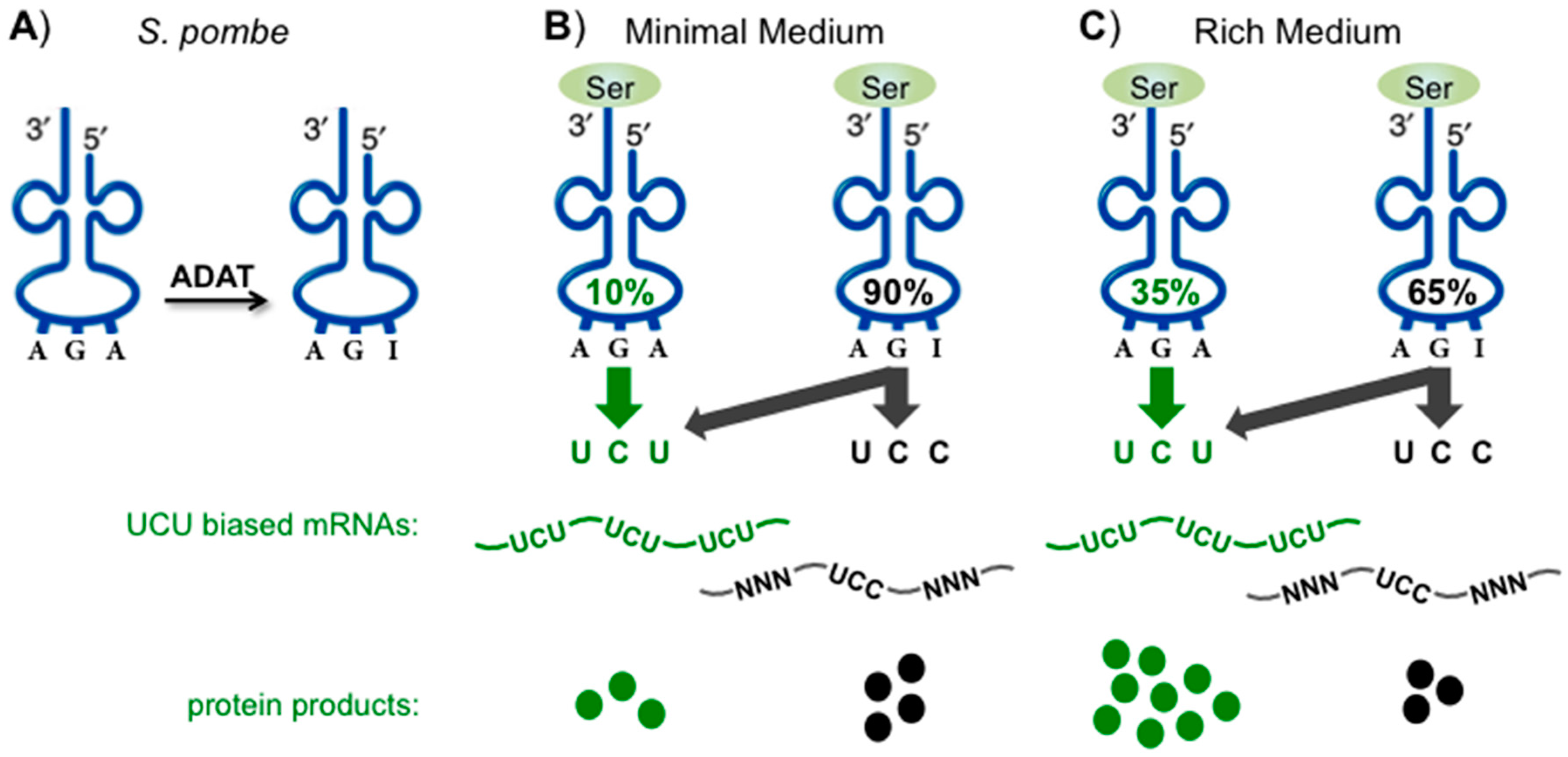
Biomolecules, Free Full-Text
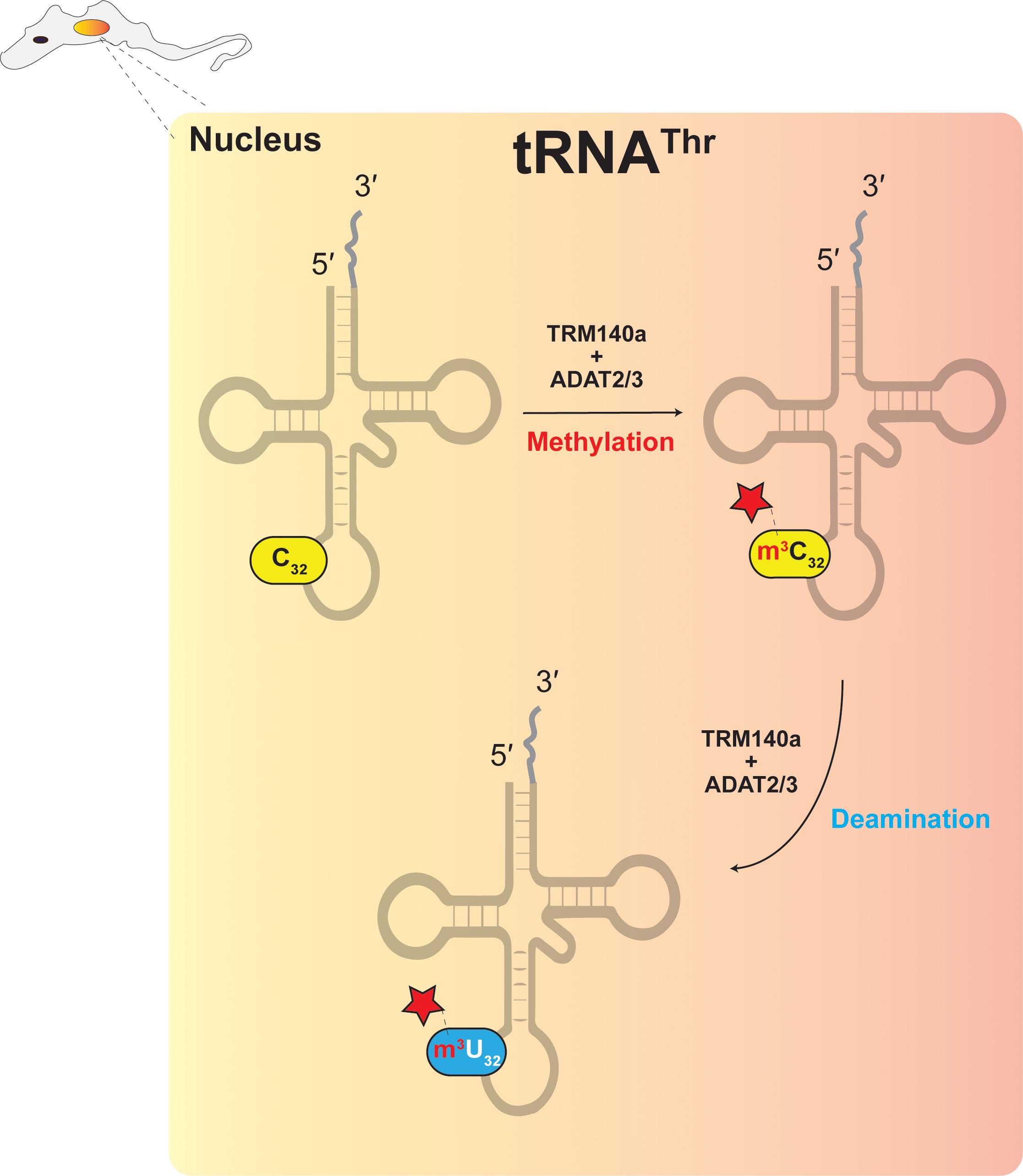
Frontiers Multi-Substrate Specificity and the Evolutionary Basis

tRNA anticodon loop signature showing the structure/sequence

PDF) Crystal Structure of tRNA Adenosine Deaminase (TadA) from

Modification of the adenosine residue into inosine at the wobble
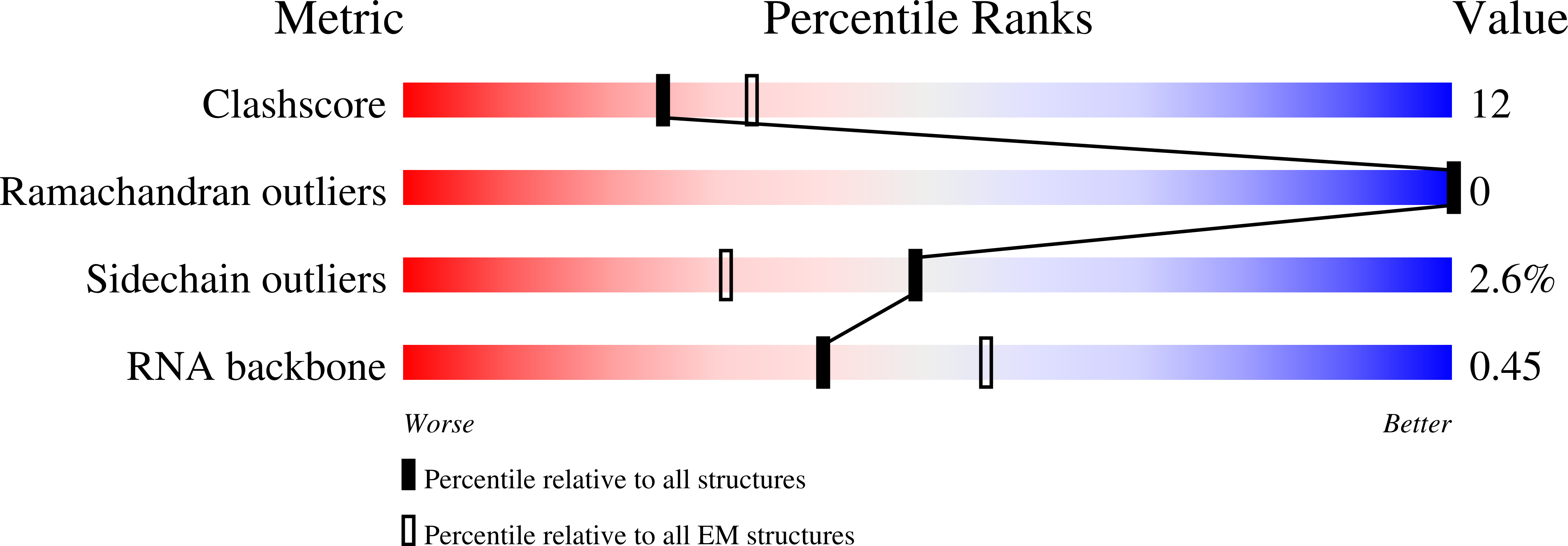
RCSB PDB - 8AW3: Cryo-EM structure of the Tb ADAT2/3 deaminase in

Structural basis of tRNA recognition by the m3C-RNA

The interaction of the tRNA anticodon-stem loop with ADAT2/3 a The

Engineered deaminases as a key component of DNA and RNA editing

Structural basis of tRNA recognition by the m3C-RNA

Phenotypic and molecular analyses of TAD2 and TAD3 RNAi mutants
from
per adult (price varies by group size)