GitHub - tgac-vumc/ACE: Absolute Copy Number Estimation using low-coverage whole genome sequencing data
By A Mystery Man Writer
Description
Absolute Copy Number Estimation using low-coverage whole genome sequencing data - tgac-vumc/ACE
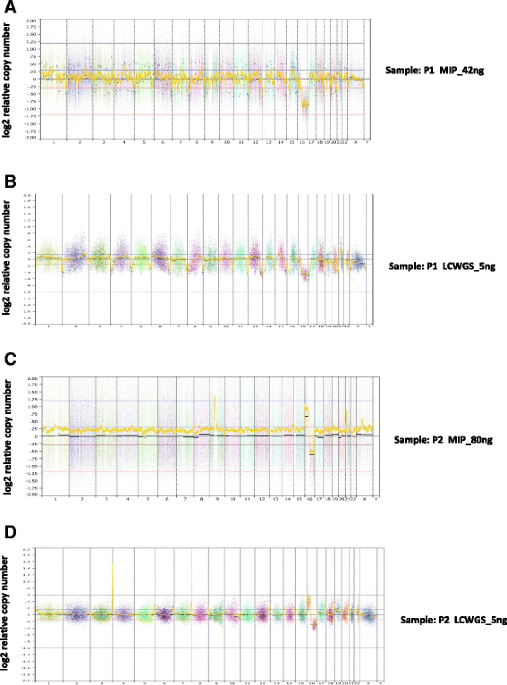
Copy number analysis by low coverage whole genome sequencing using ultra low-input DNA from formalin-fixed paraffin embedded tumor tissue, Genome Medicine

CNAplot — Software for visual inspection of chromosomal copy number alteration in cancer using juxtaposed sequencing read depth ratios and variant allele frequencies - SoftwareX
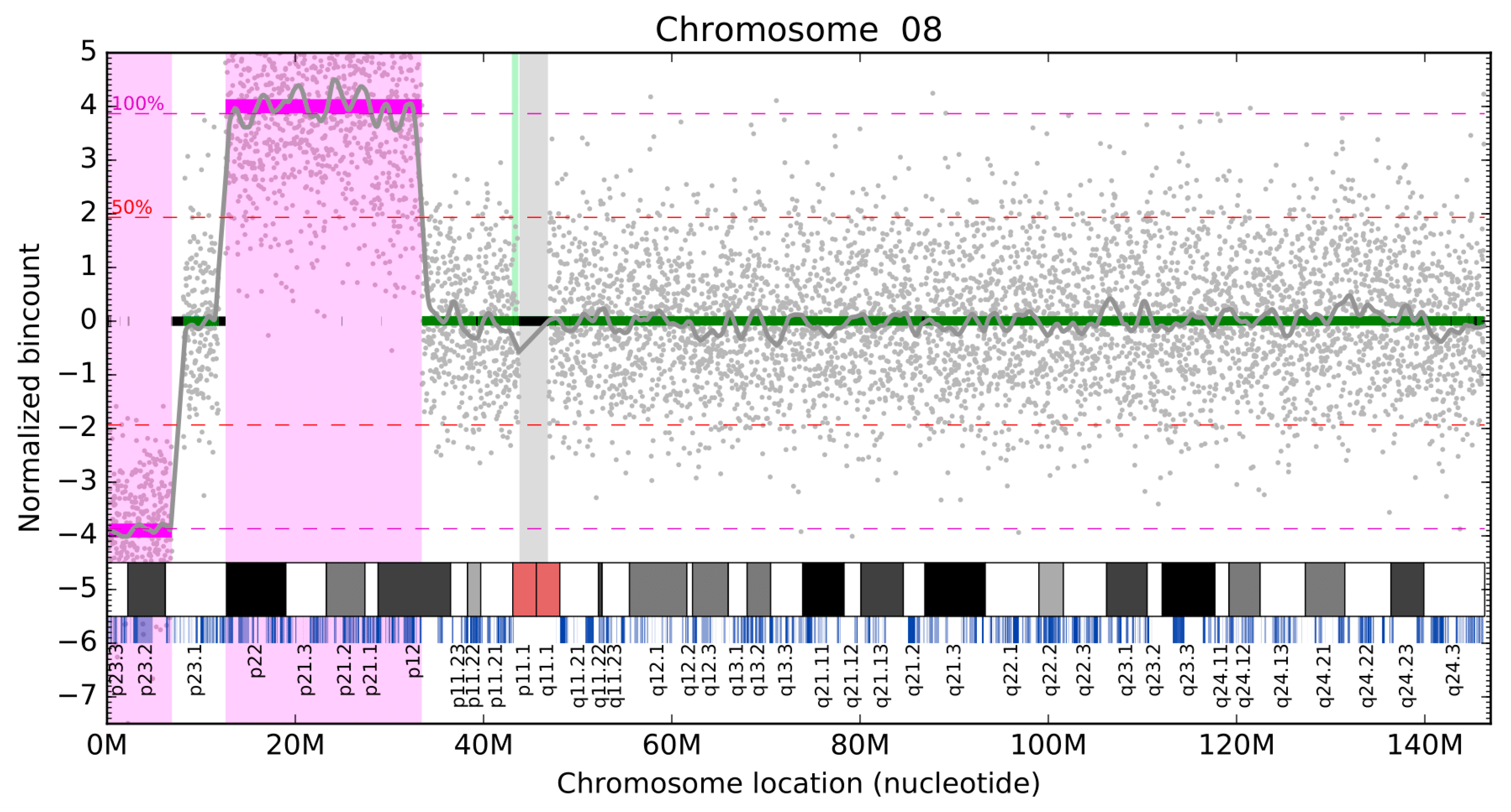
Diagnostics, Free Full-Text

Accurate quantification of copy-number aberrations and whole-genome duplications in multi-sample tumor sequencing data

GitHub - Nealelab/whole_genome_analysis_pipeline

Copy number variation detection in whole-genome sequencing data using the Bayesian information criterion
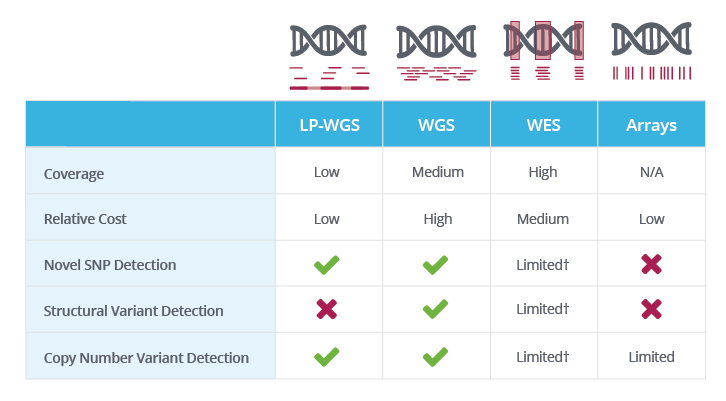
Low-Pass Whole Genome Sequencing

PDF) HBOS-CNV: A New Approach to Detect Copy Number Variations From Next-Generation Sequencing Data
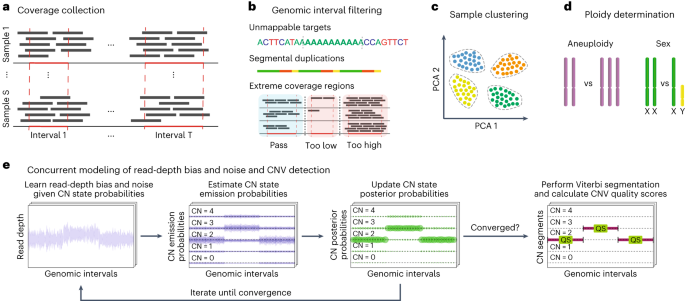
GATK-gCNV enables the discovery of rare copy number variants from exome sequencing data

Detection of Copy Number Variation using Shallow Whole Genome Sequencing Data to replace Array-Comparative Genomic Hybridization Analysis
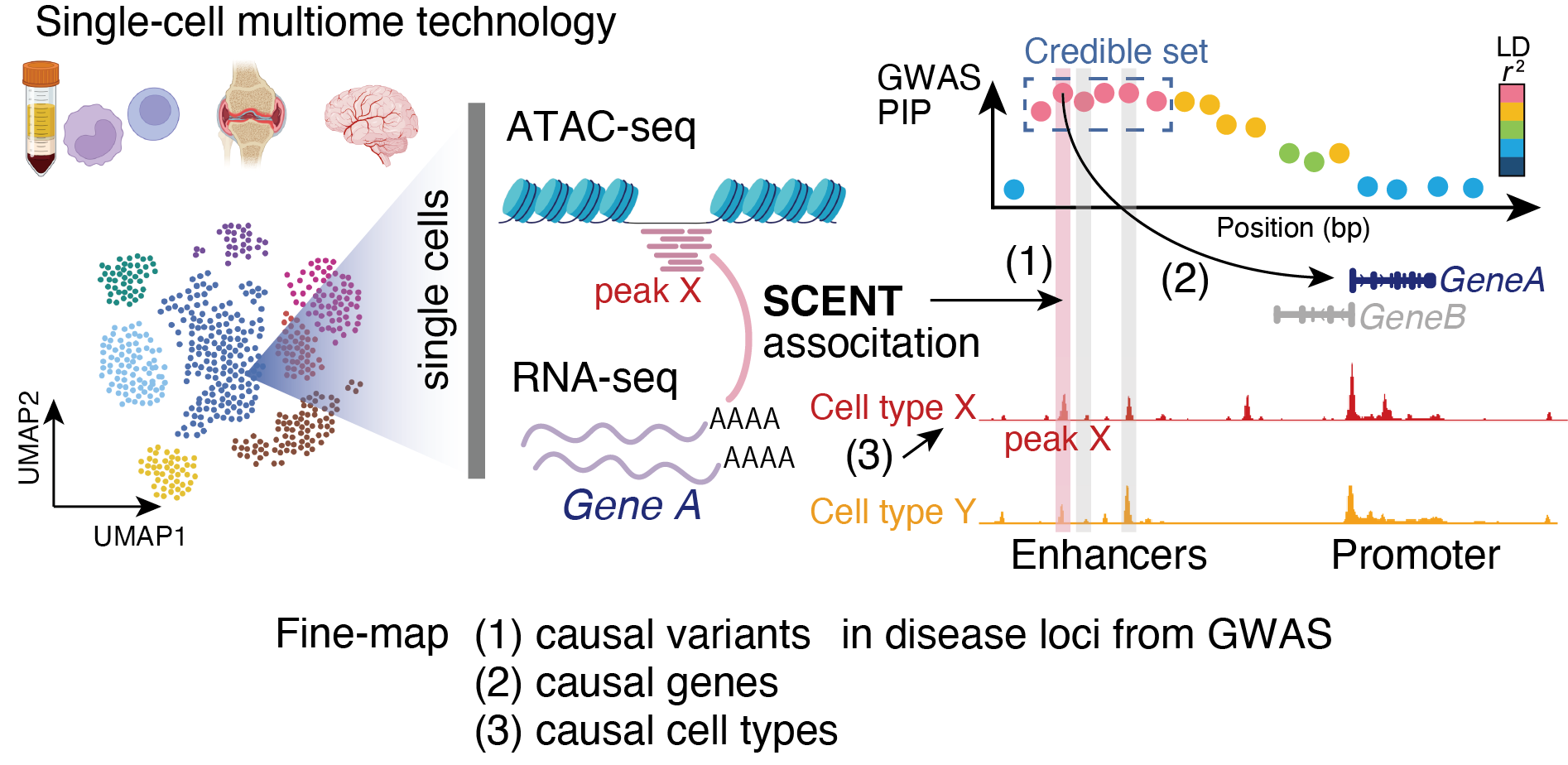
GitHub - immunogenomics/SCENT: Single-Cell ENhancer Target gene mapping using multimodal data with ATAC + RNA

Absolute copy number fitting from shallow whole genome sequencing data
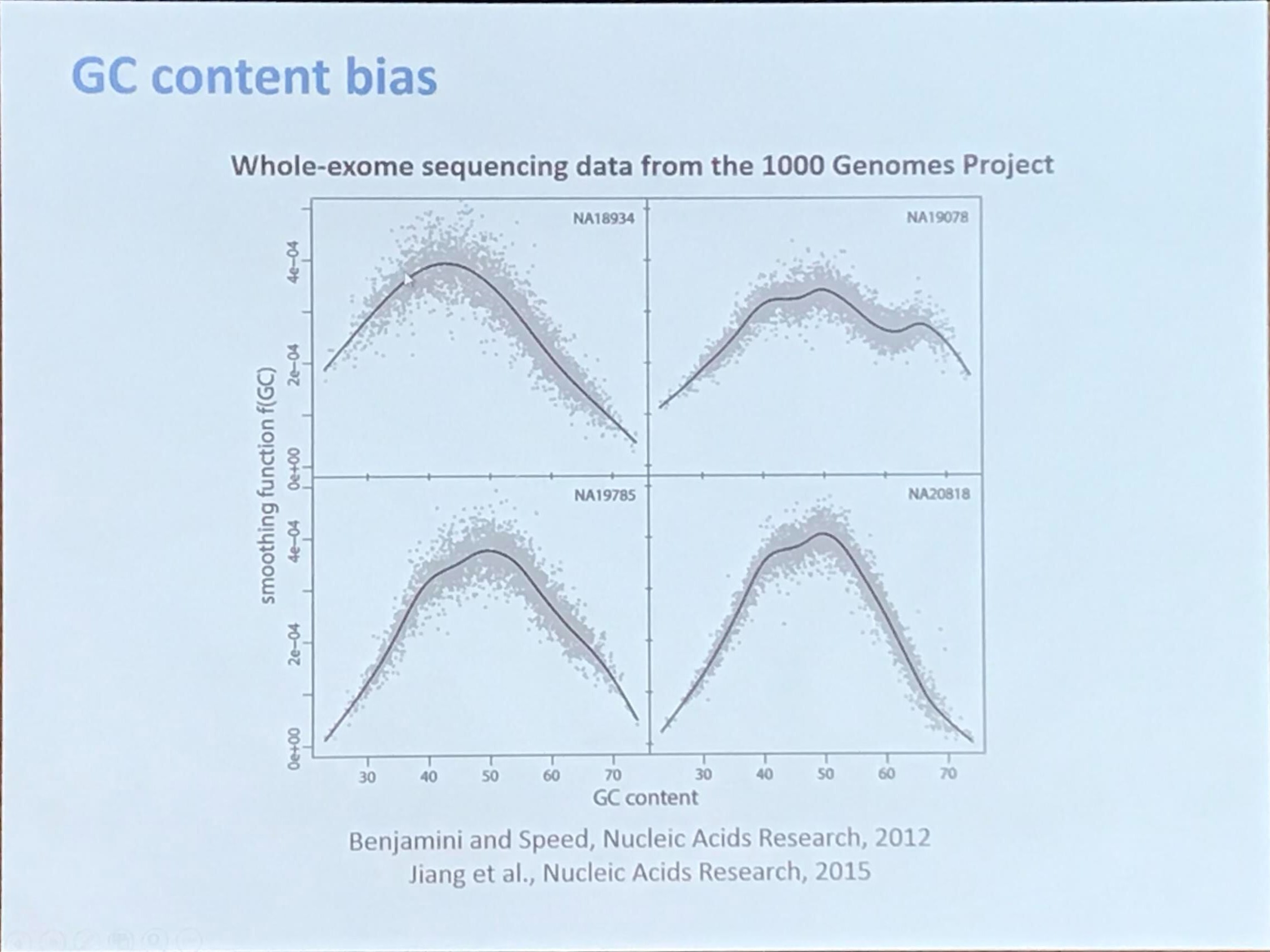
DNA copy number profiling: from bulk tissue to single cells

GATK-gCNV: A Rare Copy Number Variant Discovery Algorithm and Its Application to Exome Sequencing in the UK Biobank

PDF) WAVECNV: A New Approach for Detecting Copy Number Variation by Wavelet Clustering
from
per adult (price varies by group size)







