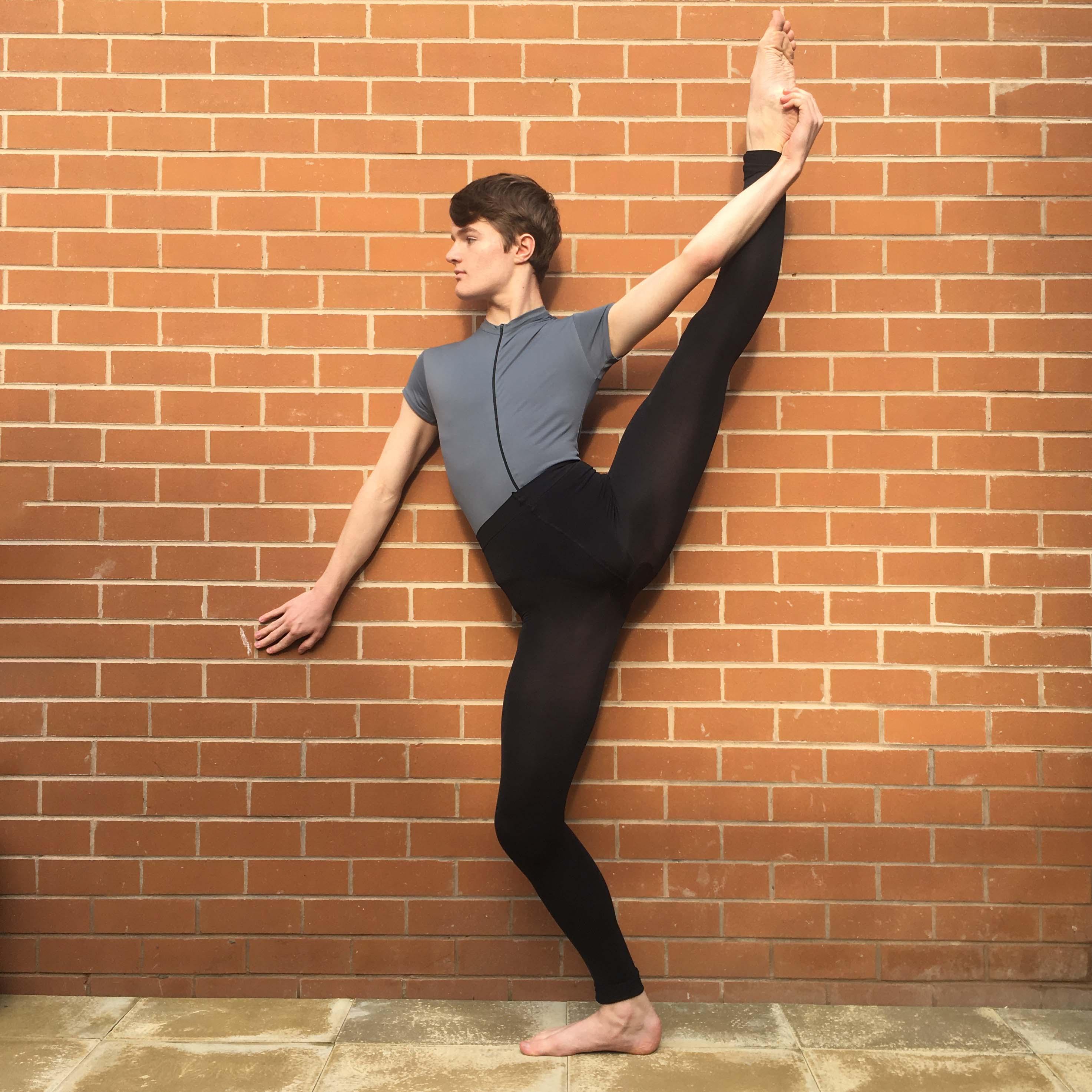
Comparison of Hi-C and Micro-C data. A Experimental methods of Hi
By A Mystery Man Writer
Description

Comparison of Hi-C and Micro-C data. A Experimental methods of Hi

PDF) Characterizing chromatin interactions of regulatory elements and nucleosome positions, using Hi-C, Micro-C, and promoter capture Micro-C

Multiscale and integrative single-cell Hi-C analysis with Higashi

bin3C: exploiting Hi-C sequencing data to accurately resolve

Comparison of computational methods for Hi-C data analysis

PDF) MoDLE: High-performance stochastic modeling of DNA loop
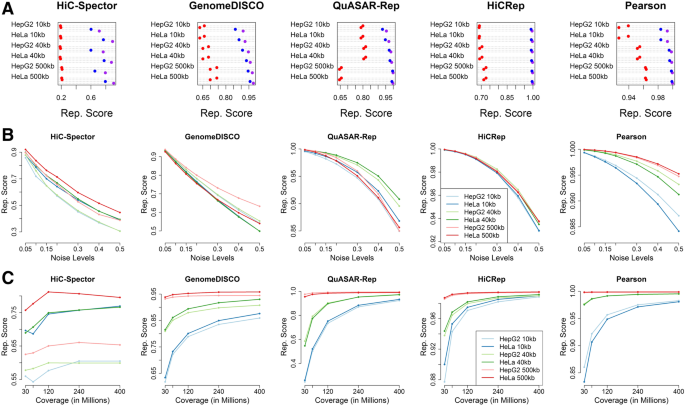
Measuring the reproducibility and quality of Hi-C data
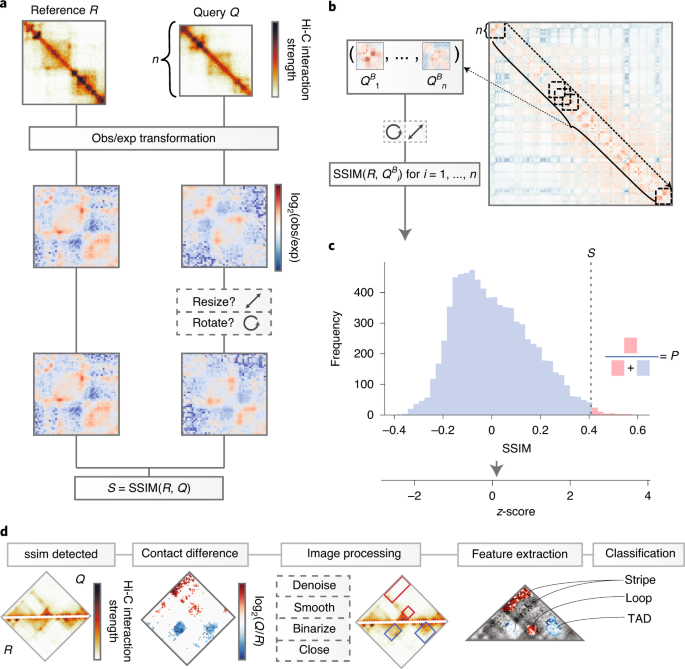
CHESS enables quantitative comparison of chromatin contact data

Comparison of the Hi-C, GAM and SPRITE methods using polymer
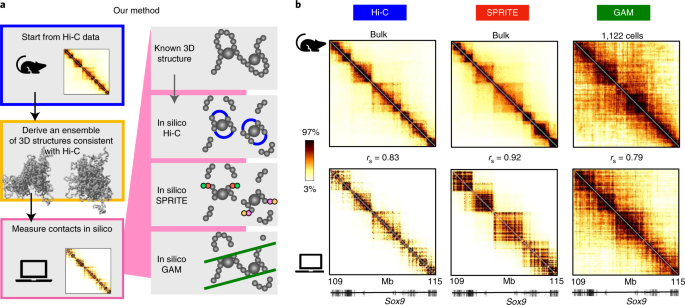
Comparison of the Hi-C, GAM and SPRITE methods using polymer
from
per adult (price varies by group size)






